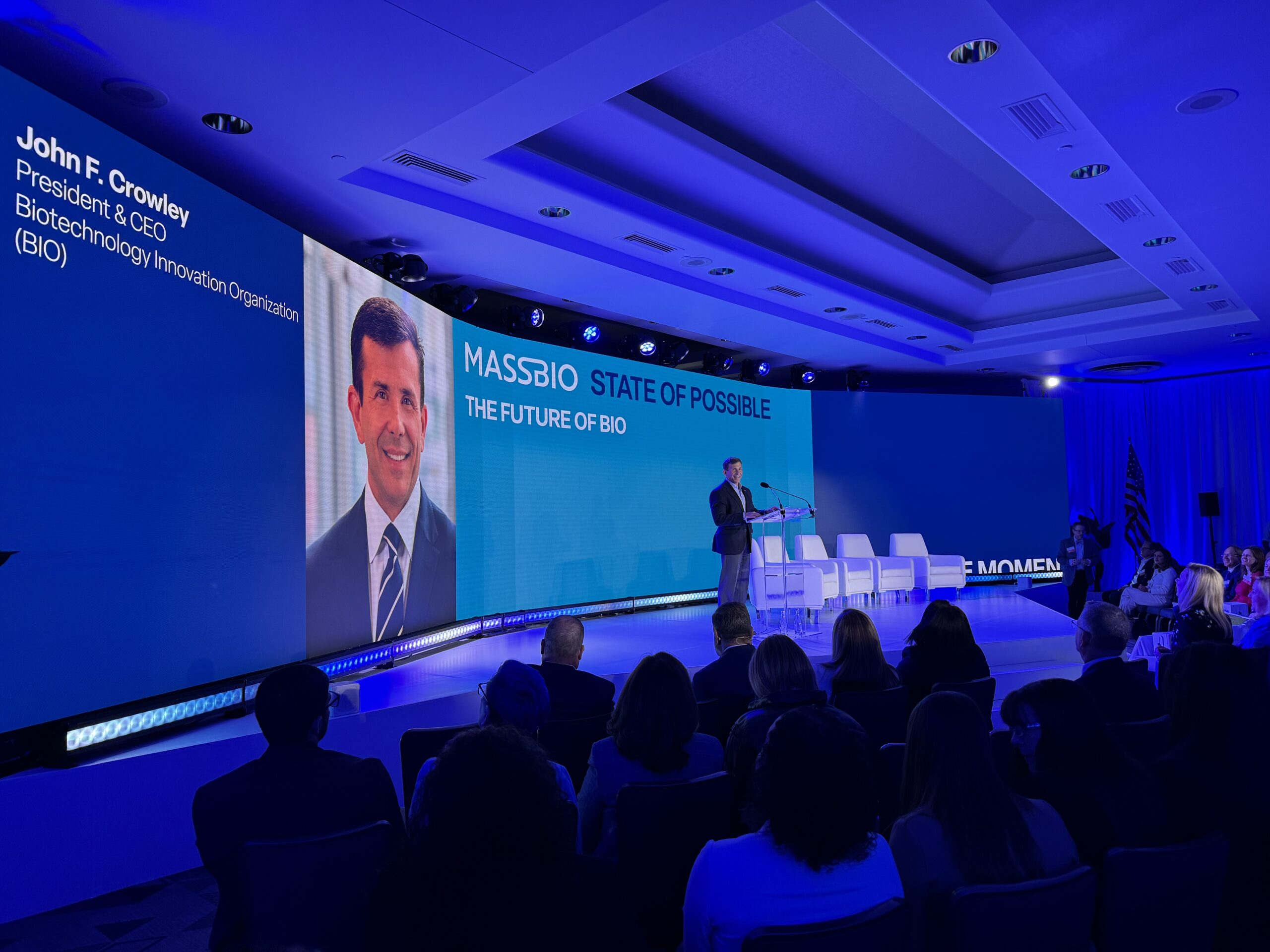
John F. Crowley: ‘We must further expand the state of possible’
On Wednesday, April 24, MassBio hosted the “State of Possible Conference,” considered “the ...
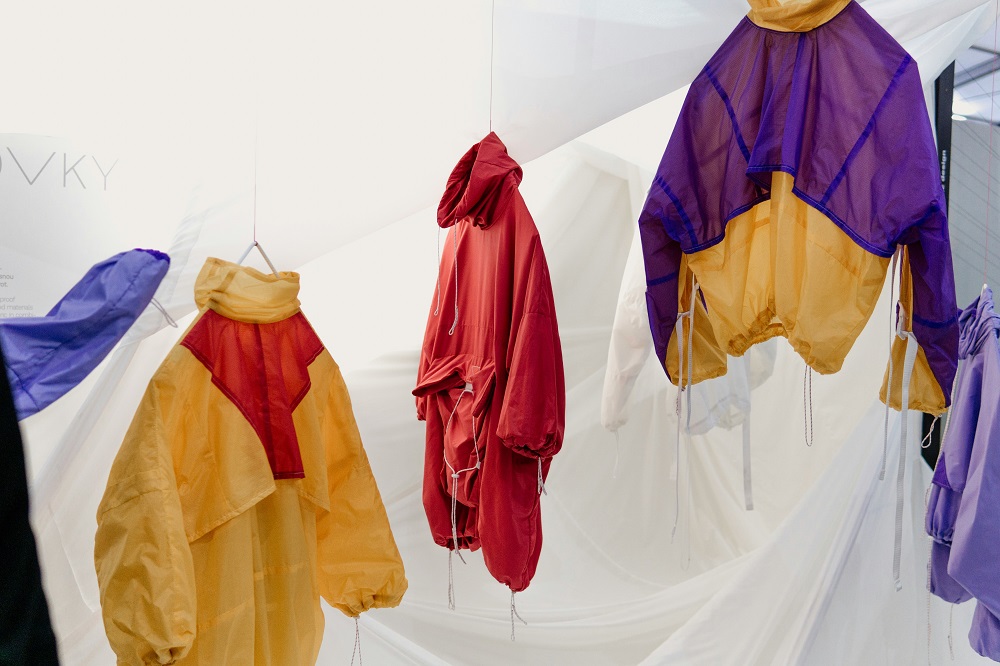
Read More LanzaTech x lululemon collab births a new sustainable fashion item
Athletic apparel retailer lululemon and biotech innovator LanzaTech have joined forces to create ...
Read More Company presentations at BIO 2024 inspire partnering
Company presentations at the BIO International Convention put innovators on stage before a ...
Read More Earth Day 2024 highlights need for plastic solutions
April 22 is Earth Day, a day that underscores the importance of conserving ...
Read More BIO Agriculture & Environment Summit brings together bipartisan policymakers, regulators
On April 18, BIO’s inaugural Agriculture & Environment Summit brought together bipartisan policymakers ...
Read More John F. Crowley: ‘Biotechnology is another word for hope’
On April 18, 2024, the Biotechnology Innovation Organization (BIO) held the inaugural Agriculture ...
Read More EDITORS' CHOICE
BIO Agriculture & Environment Summit brings together bipartisan policymakers, regulators
On April 18, BIO’s inaugural Agriculture & Environment Summit brought together bipartisan policymakers who emphasized the promise of innovation to address key challenges such as ...
April 19, 2024
Read More LATEST NEWS
BIO's View
John F. Crowley: ‘We must further expand the state of possible’
On Wednesday, April 24, MassBio hosted the “State of Possible Conference,” considered “the premier event for the Massachusetts life sciences industry.” Biotechnology Innovation Organization (BIO) ...
April 24, 2024
Biomanufacturing
LanzaTech x lululemon collab births a new sustainable fashion item
Athletic apparel retailer lululemon and biotech innovator LanzaTech have joined forces to create a sustainable clothing item in a move to reduce pollution as well ...
April 24, 2024
Agriculture
Company presentations at BIO 2024 inspire partnering
Company presentations at the BIO International Convention put innovators on stage before a small but important audience, helping them engage in the networking and partnering ...
April 23, 2024
Biomanufacturing
Earth Day 2024 highlights need for plastic solutions
April 22 is Earth Day, a day that underscores the importance of conserving our environment and advocating for greater sustainability. It’s a call to action ...
April 22, 2024
Agriculture
BIO Agriculture & Environment Summit brings together bipartisan policymakers, regulators
On April 18, BIO’s inaugural Agriculture & Environment Summit brought together bipartisan policymakers who emphasized the promise of innovation to address key challenges such as ...
April 19, 2024
HEALTH
Company presentations at BIO 2024 inspire partnering
April 23, 2024
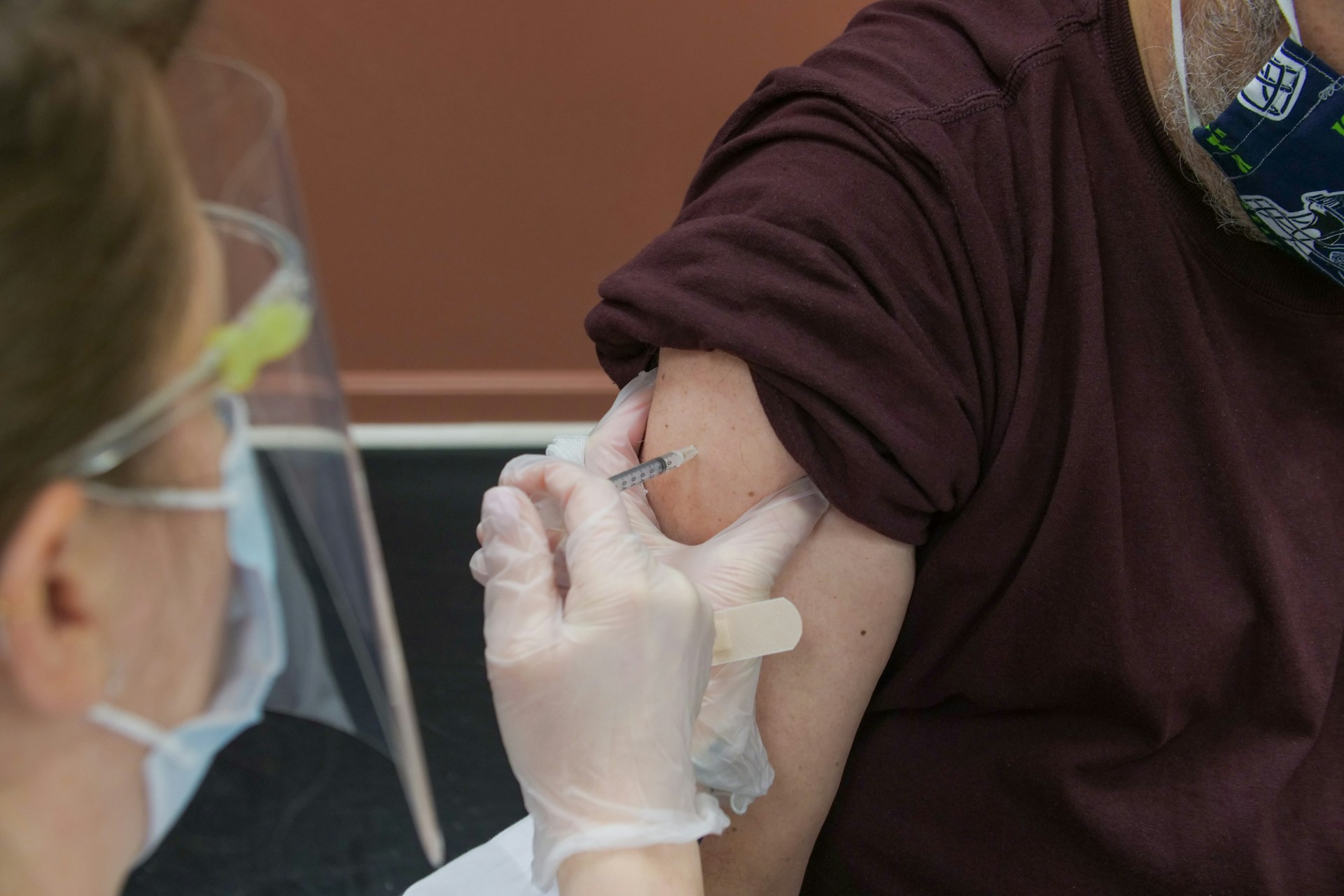
5 things to know for Primary Immunodeficiency Month
April 16, 2024
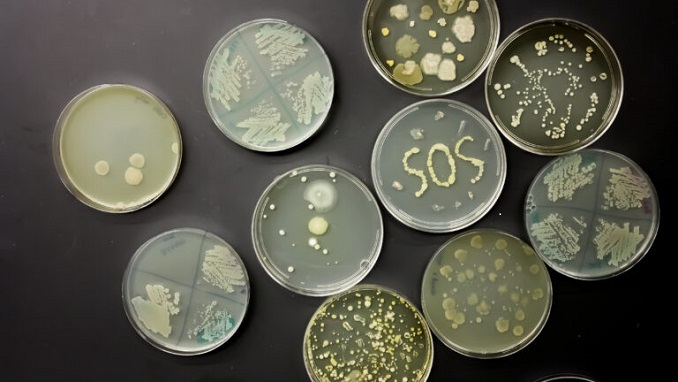
Biotech and One Health are key to controlling avian flu
April 15, 2024
Welcome, John F. Crowley!
Get to know John F. Crowley, the new President and CEO of the Biotechnology Innovation Organization (BIO), in our exclusive interview.
AGRICULTURE
Company presentations at BIO 2024 inspire partnering
April 23, 2024
John F. Crowley: ‘Biotechnology is another word for hope’
April 18, 2024
Biotech and One Health are key to controlling avian flu
April 15, 2024
Climate Change
LanzaTech x lululemon collab births a new sustainable fashion item
April 24, 2024
Athletic apparel retailer lululemon and biotech innovator LanzaTech have joined forces to create a sustainable clothing item in a move ...
Read More Earth Day 2024 highlights need for plastic solutions
April 22, 2024
John F. Crowley: ‘Biotechnology is another word for hope’
April 18, 2024
Federal Policy
State Policy
Life Sciences PA recognizes John F. Crowley for biotech leadership
April 15, 2024
On April 10, Life Sciences PA awarded Biotechnology Innovation Organization (BIO) President & CEO John F. Crowley the Hubert J.P. ...
Read More Colorado PDAB could limit patient access to medicines
March 20, 2024
New Mexico legislature passes clean fuel standard
February 19, 2024
International
5 reasons for investor optimism at BIO-Europe Spring 2024
March 29, 2024
WTO Ministerial ends, no expansion of COVID IP waiver
March 4, 2024
BIO Board member tells Congress WTO IP waivers hurt innovation
February 8, 2024
Bio's View
John F. Crowley: ‘We must further expand the state of possible’
April 24, 2024
On Wednesday, April 24, MassBio hosted the “State of Possible Conference,” considered “the premier event for the Massachusetts life sciences ...
Read More Company presentations at BIO 2024 inspire partnering
April 23, 2024
John F. Crowley: ‘Biotechnology is another word for hope’
April 18, 2024